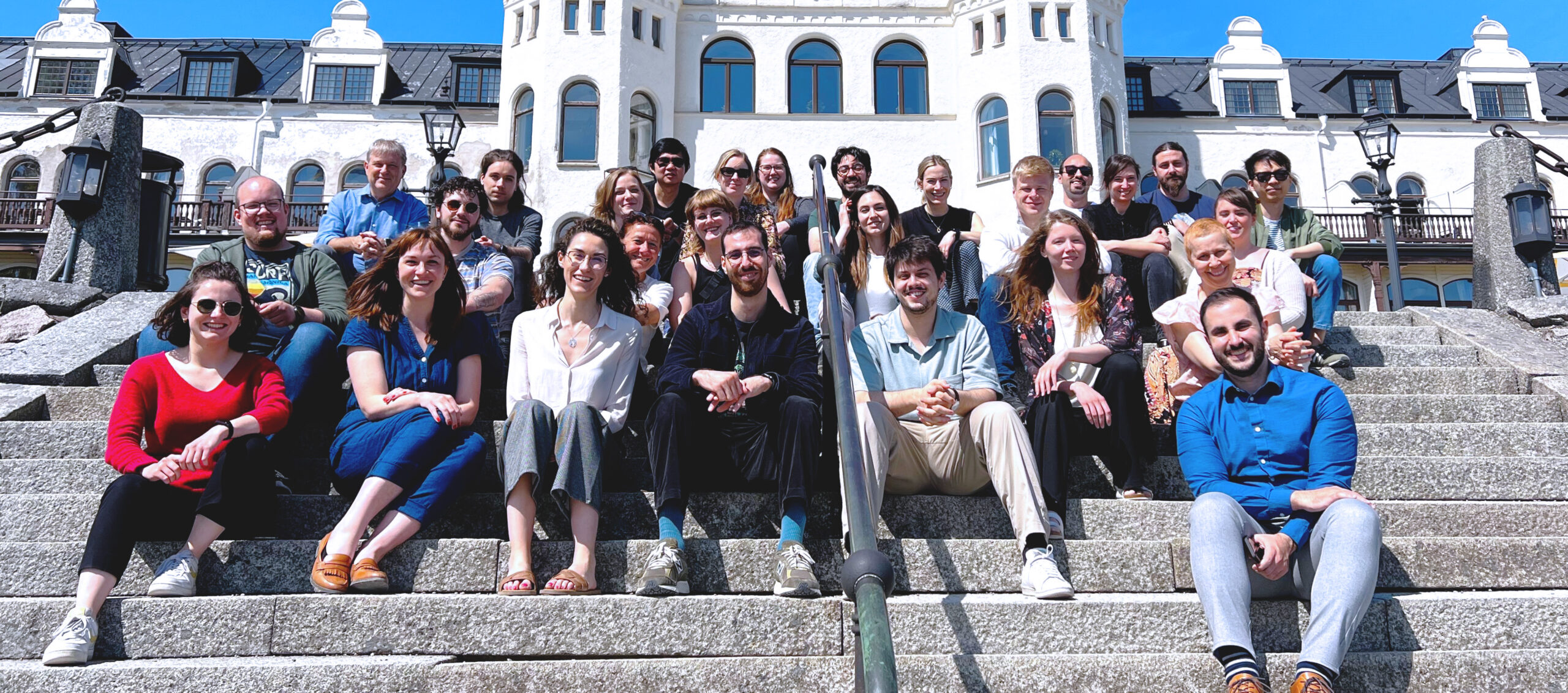

Welcome to the Spatial Research lab!

Our research focuses on developing innovative experimental methods as well as novel computational tools for spatially resolved omics in mammalian and plant tissue sections.
You can find out more about our team and our academic research activities under the Research section.

LATEST PUBLICATIONS
2025

Spatiotemporal gene expression and cellular dynamics of the developing human heart
Enikő Lázár, Raphaël Mauron, Žaneta Andrusivová, Julia Foyer, Mengxiao He, Ludvig Larsson, Nick Shakari, Sergio Marco Salas, Christophe Avenel, Sanem Sariyar, Jan Niklas Hansen, Marco Vicari, Paulo Czarnewski, Emelie Braun, Xiaofei Li, Olaf Bergmann, Christer Sylvén, Emma Lundberg, Sten Linnarsson, Mats Nilsson, Erik Sundström, Igor Adameyko, Joakim Lundeberg
Nature Genetics, 29 October 2025

Spatial architecture of development and disease
Enikő Lázár, Joakim Lundeberg
Nature Review Genetics, 30 September 2025

Computational pathology annotation enhances the resolution and interpretation of breast cancer spatial transcriptomics data
Li T, Yang Q, Acs B, Sifakis E, Toosi H, Engblom C, Thrane K, Lin Q, Mold J, Sun W, Boyaci C, Steen S, Frisén J, Largergren J, Lundeberg J, Chen X, Hartman J
Nature Precision Oncology, 9 September 2025

adiposetissue.org: A knowledge portal integrating clinical and experimental data from human adipose tissue
Zhong J, Zareifi D, Weinbrenner S, Hansen M, Klingelhuber F, Nono Nankam PA, Frendo-Cumbo S, Bhalla N, Cordeddu L, de Castro Barbosa T, Arner P, Dahlman I, Muniandy M, Heinonen S, Pietiläinen KH, Hoffmann A, Ghosh A, John D, Tönjes A, Ståhl PL, Böttcher Y, Keller M, Kovacs P, Kerr AG, Langin D, Wolfrum C, Blüher M, Krahmer N, Massier L, Mejhert N, Rydén M
Cell Metabolism, 14 February 2025
Preprints 2025
Spatially resolved integrative analysis of transcriptomic and metabolomic changes in tissue injury studies
Williams EC, Franzén L, Olsson Lindvall M, Hamm G, Oag S, Majumder MM, Denholm J, Hamidinekoo A, Escudero Morlanes J, Vicari M, Lundeberg J, Setyo L, Zakirov A, Hornberg JJ, Stamou M, Ståhl PL, Ollerstam A, Tan J, Mohorianu I
bioRxiv, 2 March 2025
2024

scTrends: A living review of commercial single-cell and spatial ‘omic technologies
De Jonghe J, Opzoomer JW, Vilas-Zornoza A, Nilges BS, Crane P, Vicari M, Lee H, Lara-Astiaso D, Gross T, Morf J, Schneider K, Cudini J, Ramos-Mucci L, Mooijman D, Tiklová K, Salas SM, Langseth CM, Kashikar ND; scTrends Consortium; Schapiro D, Lundeberg J, Nilsson M, Shalek AK, Cribbs AP, Taylor-King JP
Cell Genomics, 11 December 2024

Spatial transcriptomics reveals substantial heterogeneity in triple-negative breast cancer with potential clinical implications
Wang X, Venet D, Lifrange F, Larsimont D, Rediti M, Stenbeck L, Dupont F, Rouas G, Garcia AJ, Craciun L, Buisseret L, Ignatiadis M, Carausu M, Bhalla N, Masarapu Y, Villacampa EG, Franzén L, Saarenpää S, Kvastad L, Thrane K, Lundeberg J, Rothé F, Sotiriou C.
Nature Communications, 26 November 2024

Standalone single- and bi-layered human skin 3D models supported by recombinant silk feature native spatial organization
Gkouma S, Bhalla N, Frapard S, Jönsson A, Gürbüz H, Dogan AA, Giacomello S, Duvfa M, Ståhl PL, Widhe M, Hedhammar M
Biofabrication, 5 November 2024
Contact
Joakim Lundeberg
Professor

![]()
Science for Life Laboratory
KTH, School of Engineering Sciences in Chemistry, Biotechnology and Health, Dept. of Gene Technology
Tomtebodavägen 23 A
171 65 Solna, Sweden![]()
joakim.lundeberg@scilifelab.se
Stefania Giacomello
Associate Professor, Ph.D

![]()
Science for Life Laboratory
KTH, School of Engineering Sciences in Chemistry, Biotechnology and Health, Dept. of Gene Technology
Tomtebodavägen 23 A
171 65 Solna, Sweden![]()
stefania.giacomello@scilifelab.se
Patrik Ståhl
Professor, Ph.D

![]()
Science for Life Laboratory
KTH, School of Engineering Sciences in Chemistry, Biotechnology and Health, Dept. of Gene Technology
Tomtebodavägen 23 A
171 65 Solna, Sweden![]()
patrik.stahl@scilifelab.se
Jonas Frisén
Professor

![]()
Karolinska Institutet,
Biomedicum B6,
Solnavägen 9,
171 65 Solna, Sweden![]()
jonas.frisen@ki.se

