The Spatial Research Lab
We are a big bunch of researchers based in Stockholm, Sweden, all focused on looking at the spatial gene expression of tissues. The groups led by PIs Joakim Lundeberg, Patrik Ståhl, and Stefania Giacomello (KTH Royal Institute of Technology) work closely together and we have our lab at the Science for Life Laboratory (SciLifeLab). We are also well connected to the group led by Jonas Frisén, affiliated and based on the Karolinska Institute campus.
Our research is mainly focused on developing and exploring spatially resolved gene expression tools and their applications. Many of our projects revolves on building large scale interdisciplinary molecular atlases, including human development (see HDCA), though we also conduct research looking at tumour heterogeneity, various human diseases where cell type and location plays a vital role, the effect of spaceflight, and tissue expression in plant species.

The Spatial Transcriptomics / Visium technology
Spatial Transcriptomics (ST) is a method that allows visualization and quantitative analysis of the transcriptome in individual tissue sections. By placing histological sections on glass slides with arrayed oligonucleotides containing positional barcodes, it is possible to generate high quality cDNA libraries with precise positional information for RNA-seq. This provides transcriptome data in a versatile format for bioinformatics analyses of gene expression within the tissue context, which will be valuable in both research and diagnostics. The invention was originally published 2016 in Science by Ståhl and Salmén et al.
Prof Joakim Lundeberg (KTH Royal Institute of Technology) and Prof Jonas Frisén (Karolinska Institutet) received a key initial support from the Knut and Alice Wallenberg Foundation in 2012 to develop and use the Spatial Transcriptomics technology for analysis and discovery of transcriptional patterns in tissue, with a focus on the brain. The method has received increasing attention and is currently the basis of several national and international collaborations. The research is predominantly done at Science for Life Laboratory, Stockholm.
In 2019, the technology was acquired by 10x Genomics and launched under the product name Visium. The Visium platform was further developed to have a smoother and faster workflow as well as producing a larger amount of data per tissue section by improving the spatial resolution. Our lab is in a continuous collaboration with 10x Genomics and leading the scientific progress made within the area of spatially resolved transcriptomics.
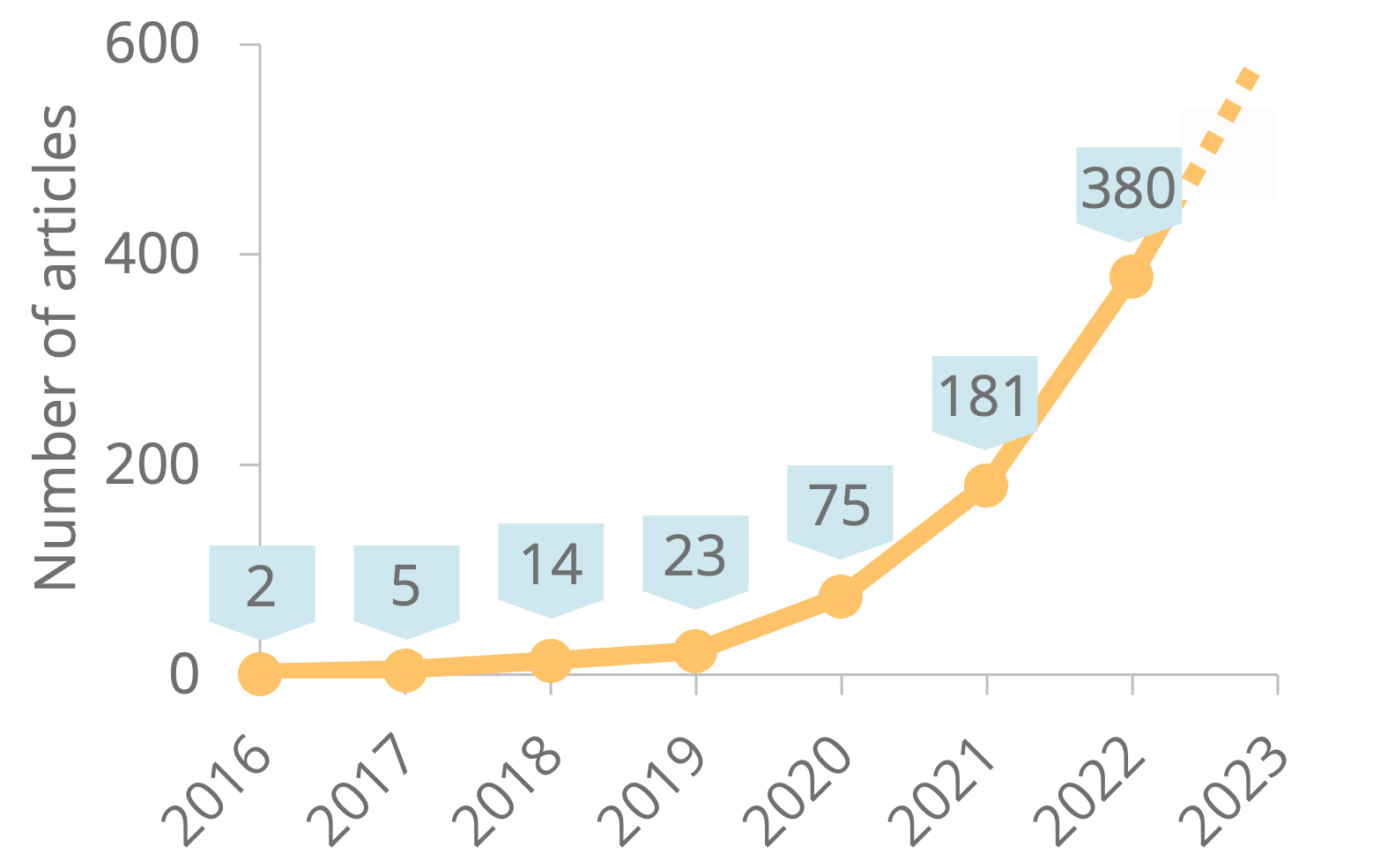
The field of spatially resolved transcriptomics and omics have exploded in the recent years, and Nature Methods announced the technology “Method of the year 2020”. Ludvig Larsson, Jonas Frisén, and Joakim Lundeberg were given the honour of publishing their commentary “Spatially resolved transcriptomics adds a new dimension to genomics“ as experts on this subject.

Image: Ludvig Larsson, Natalie Stakenborg, Joakim Lundeberg and Guy Boeckxstaens
